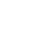
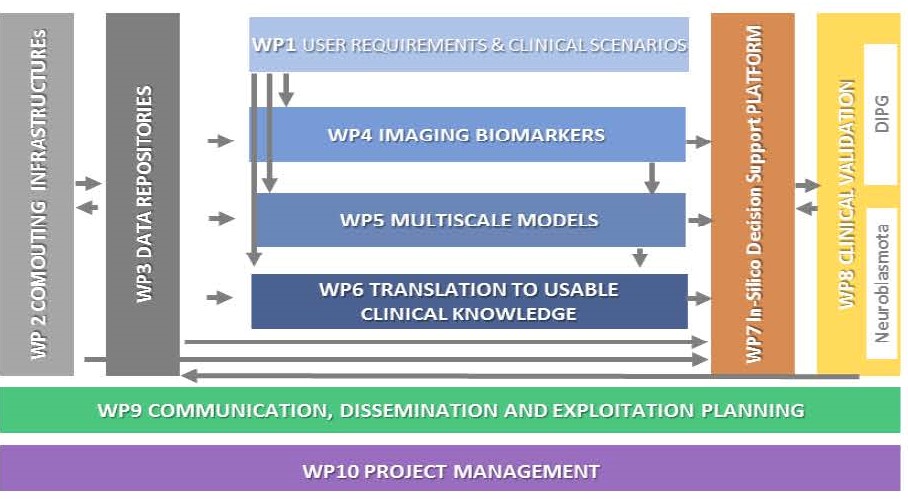
This project has a 48-months duration and is structured in 10 Work Packages (WP):
WP1 – Clinical scenarios & use requirements
WP1’s successfully reached objective was to design a decision support system for cancer management with advanced functionality and usability under a user-centric approach.
CCRI managed the input and coordination between the different data repositories and the many different ways of accessing retrospective clinical, imaging and biological data from clinical study registries and hospitals.
Before uploading and using data in the PRIMAGE Platform, the consortium decided to pseudonymise data via EUPID to ensure data handling according to data protection rules, an essential condition for all WPs and their integrated tasks.
| WP1 – Deliverables | |
| D1.1 | Users´ requirements and use scenarios (confidential) |
| D1.2 | Platform architecture, interconnectivities and standards (confidential) |
WP2 – Computing infrastructures
A cloud and an HPC infrastructure supports the PRIMAGE platform and provides storage and processing resources for the project data lakes, replicated among providers to guarantee high availability and robustness.
This infrastructure uses shared European infrastructures (PLGrid, EGI) and services from the European Open Science (EOSC) Marketplace. It comprises open-source components, and its code aims for deployment on different providers with minimal user interaction. Although specially tailored for medical imaging data analytics, the infrastructure is agnostic to the cloud provider and can fit other problems related to data analytics.
| WP2 – Deliverables | |
| D2.1 | Description of available HPC resources, UI/API components access procedures (confidential) |
| D2.2 | Description of Hybrid Open Datacloud and processing middleware architecture (confidential) |
| D2.3 | Deployment of intermediate version of hybrid open cloud and processing middleware (confidential) |
WP3 – Data Repositories
WP3 aims to facilitate the extraction of usable and meaningful imaging biomarkers from tissue and liquid biopsies and reuse them in imaging biobanks.
The extraction will establish imaging data collection and annotation recommendations, streamline the process, and facilitate its exploitation and valorisation as high-quality training data sets. The proposed methodologies and protocols for data management will lower legal, administrative, and technical barriers, thus promoting an Open Science culture that embeds imaging biobanks into wider biobanks networks and facilitates data cross-linking with other biorepositories.
| WP3 – Deliverables | |
| D3.1 | Definition of data requirements, e-forms, QC procedures and signed authorisations (Public) |
| D3.2 | Implementation of PRIMAGE database and mechanisms for archiving and storing imaging data, linkable to clinical data (confidential) |
WP4 – Imaging Biomarkers
WP4 aims to deliver a standardised format of imaging and molecular biomarkers of patients with neuroblastoma and DIPG to address particular challenges in paediatric cancer imaging. To address this challenge, partners in WP4 established a common framework to assess extracted quantitative imaging descriptors’ data quality using different image analysis methods and tools.
The unsupervised learning algorithms helped: discover radiomics signatures; assess the different types of processed data; train a deep convolutional neural network to automatically segment the neuroblastic and DIPG tumours in unseen images. This process led to the development of AI models that predict different clinical outcomes by integrating molecular, biological and genomic biomarkers with imaging and clinical data.
| WP4 – Deliverables | |
| D4.1 | Generation of radiomic data from retrospective sets (Confidential) |
WP5 – Multiscale Models
WP5 focuses on developing a multiscale computational model of paediatric solid tumour growth for Neuroblastoma and Diffuse Intrinsic Pontine Glioma to simulate different clinical treatments that predict the tumour’s temporal evolution.
- Provide quantitative data for clinical use;
- Valuable information for selecting the optimal chemotherapeutic treatment for individual patients;
- Integrate genetic information to the cellular and whole tumour scale informed by imaging data, respectively;
- Improve the understanding of tumour growth mechanisms from existing data
The development of in vitro experiments of tumour spheroid growth will further refine the multiscale model and simulate the complexity of the underlying phenomena.
WP6 – Translation to Usable Clinical Knowledge
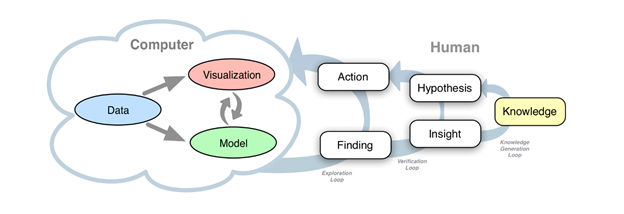
WP6 will combine retrospective patient data (WP3) with novel imaging biomarkers (WP4) and innovative in-silico models (WP5), allowing medical doctors and other domain experts to gain insight into the complex connections between these sources of information. WP6 will create a visual interactive platform for exploratory analysis and sense-making, integrating modern AI techniques with domain users’ expertise. WP6 will focus on four use cases:
- UC1: Visual Analysis of Similar Patients
- UC2: Visual Analysis of Outcome Predictions
- UC3: Visual Analysis of Measurement Correlations
- UC4: Visual Analysis of Outcome Sequences
| WP6 – Deliverables | |
| D6.1 | Requirement analysis for the design of the visual analytics system (Confidential) |
WP7 – Platform Integration
WP7 will focus on the architecture and integration of the PRIMAGE platform, which will offer predictive tools to assist disease management from diagnosis to prognosis and personalised treatment to assist clinicians in predicting clinical points of great interest. WP7 partners will implement advanced visualisation tools, visualising uncertainty and prediction reliability of in-silico models to facilitate clinical decision-making. Exhaustive lab testing will lead to delivering a beta version of the PRIMAGE platform ready to be validated in the context of paediatric cancers NB and DIPG.
WP8 – Clinical Validation
Partners in WP8 will validate the PRIMAGE platform performance with clinical paediatric oncologists (from hospitals in Spain, Germany and Austria) in prospective non-interventional studies, demonstrating the PRIMAGE platform as a helpful tool in routine clinical practice. WP8 will run during the whole project to recruit the expected cohort and use the prospective data only for platform validation during year 4.
| WP8 – Deliverables | |
| D8.1 | Readiness to start recruitment for clinical tests (Public) |
WP9 – Communication, Dissemination & Exploitation
WP9 encompasses the communication, dissemination and exploitation needs of the PRIMAGE Project. Together with the partners, SIOP Europe will develop the dissemination, communication, and networking tools and actions to network PRIMAGE’s results with relevant national and international projects, initiatives, and networks. MATICAL, together with the partners, will oversee innovation management, including ownership, access rights, decision-making procedures, publications, IP management, and business, sustainability, and exploitation planning.
| WP9 – Deliverables | |
| D9.1 | Data Management Plan (Public) |
| D9.2 | First year plan for dissemination and communication activities, incl website and dissemination toolkit 1 (Public) |
| D9.3 | Record of dissemination and communication activities in year 1 and plan for year 2, incl. Dissemination toolkit 2 (Public) |
| D9.4 | Initial exploitation plan (Confidential) |
WP10 – Project Management
WP10 comprises the day to day running of the project, ensuring that the work programme adheres to its schedule, fulfils its objectives, and the project meets the needs and expectations of all the partners. Together with the Project Manager, the Coordinator will steer the evolution of the project by closely monitoring the work performed by each partner. Both will assume responsibility for the correct implementation of project management procedures and meeting all deadlines and obligations.
| WP10 – Deliverables | |
| D10.1 | Project management tools (Confidential) |
| D10.2 | Ethics review plan (Public) |



 on Twitter
on Twitter on LinkedIn
on LinkedIn